The project team realised this honours project at the Chair of Scientific Computing (TUM) in cooperation with General Electric.
Magnetic Resonance Imaging (MRI) is a technique which is widely used in medical research and clinical applications for imaging of soft tissues. It is based on effects of the internal magnetization of atoms and yields cross-section slices of the body. Both hard- and software of MRI systems are continuously improved to get better image quality or to speed up the image formation process. While MRI is based on well-understood principles, building an actual machine presents numerous difficulties, including high cost. For this reason, simulations can be used to predict the properties of newly designed systems. Consequently, we are presenting a project which involves creating a self-contained, portable MRI simulation tool, which is to be used on various machines.
The simulation is performed slice by slice, using the isochromat summation method, where the investigated object is represented by its spin density and relaxation parameters. Further inputs are the pulse sequence and the user defined parameters. The algorithm then solves the Bloch equation for the given specifications and generates the final image of the slice by taking the Fourier transform of the resulting k-space data.
In order to obtain a 3D object, multiple cross-section slices of the object need to be simulated, where it can be assumed that the slices are independent of each other. As the calculations are computationally very intensive and therefore time-consuming, it is critical to improve their speed to achieve better resolution and enable simulating larger objects. A natural approach to this problem is parallelization, either on the CPUs or GPUs. In this project, these methods are combined by porting an existing serial isochromat summation code to the FRAVE, a multi-processor multi-GPU system at the department of computer science at TUM.
FRAVE stands for “fully reconfigurable cave environment” and, apart from multiple computers and computing GPUs, it consists of 10 screens that can be configured to a virtual reality environment and give a great possibility to visualize large 3D objects.

Illustration 1: FRAVE setup at TUM
In order to achieve portability, the application was written using cross-platform libraries which can be compiled for both Windows and Linux. This GPU-accelerated simulator is written in C++ with CUDA. When given a virtual 3-D object, the simulation processes it on a slice-by-slice basis, dividing the slices among the compute nodes at FRAVE, where at most 3 compute nodes are used at the time. The communication between different compute nodes was performed using the MPI library. The results can be visualized either on a computer or on the FRAVE using 6 screens. OpenGL is used for volume rendering by ray-casting and a slice-by-slice walk-through enables seeing 2D slices separately. For the visualization on the FRAVE, the Equalizer library is used for parallel rendering. Specifying the input parameters can be done through the graphical user interface which was created using Qt.
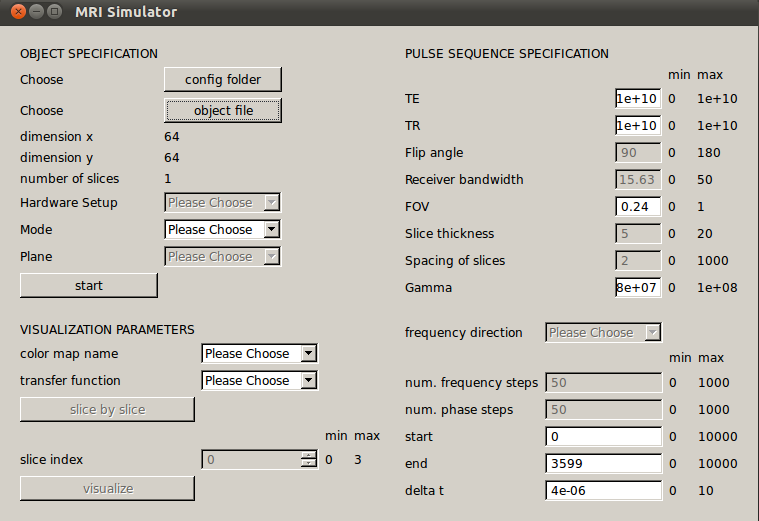
Illustration 2: Graphical User Interface
In comparison to the initial sequential implementation, we achieved a speed-up of roughly 8 times per slice. The speed of the simulation of the whole 3D object increases linearly with the number of compute nodes used for the simulation.
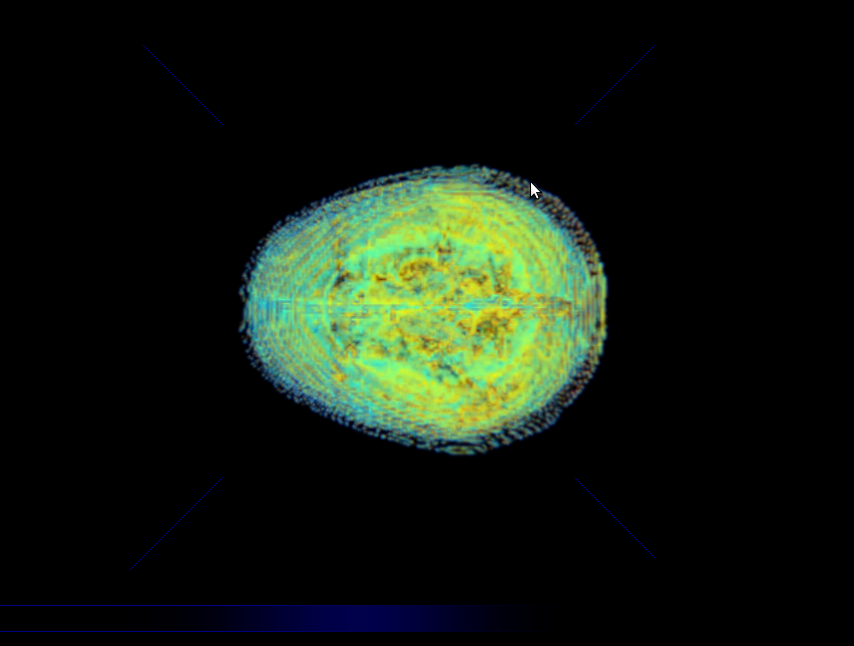
Illustration 3: Simulation Results for a Brain Model
